Principles of Biochemistry/Nucleic acid II: RNA and its nucleotides
| This page needs to be de-wikified. Books should use wikilinks rather sparsely, and only to reference technical or esoteric terms that are critical to understanding the content. Most if not all wikilinks should simply be removed. Please remove {{dewikify}} after the page is dewikified. |
Ribonucleic acid (RNA) is one of the three major macromolecules (along with DNA and proteins) that are essential for all known forms of life.
Like DNA, RNA is made up of a long chain of components called nucleotides. Each nucleotide consists of a nucleobase (sometimes called a nitrogenous base), a ribose sugar, and a phosphate group. The sequence of nucleotides allows RNA to encode genetic information. For example, some viruses use RNA instead of DNA as their genetic material, and all organisms use messenger RNA (mRNA) to carry the genetic information that directs the synthesis of proteins. Like proteins, some RNA molecules play an active role in cells by catalyzing biological reactions, controlling gene expression, or sensing and communicating responses to cellular signals. One of these active processes is protein synthesis, a universal function whereby mRNA molecules direct the assembly of proteins on ribosomes. This process uses transfer RNA (tRNA) molecules to deliver amino acids to the ribosome, where ribosomal RNA (rRNA) links amino acids together to form proteins. The chemical structure of RNA is very similar to that of DNA, with two differences--(a) RNA contains the sugar ribose while DNA contains the slightly different sugar deoxyribose (a type of ribose that lacks one oxygen atom), and (b) RNA has the nucleobase uracil while DNA contains thymine (uracil and thymine have similar base-pairing properties). Unlike DNA, most RNA molecules are single-stranded. Single-stranded RNA molecules adopt very complex three-dimensional structures, since they are not restricted to the repetitive double-helical form of double-stranded DNA. RNA is made within living cells by RNA polymerases, enzymes that act to copy a DNA or RNA template into a new RNA strand through processes known as transcription or RNA replication, respectively[1].
Structure of RNA
[edit | edit source]

Each nucleotide in RNA contains a ribose sugar, with carbons numbered 1' through 5'. A base is attached to the 1' position, in general, adenine (A), cytosine (C), guanine (G), or uracil (U). Adenine and guanine are purines, cytosine, and uracil are pyrimidines. A phosphate group is attached to the 3' position of one ribose and the 5' position of the next. The phosphate groups have a negative charge each at physiological pH, making RNA a charged molecule (polyanion). The bases may form hydrogen bonds between cytosine and guanine, between adenine and uracil and between guanine and uracil.However, other interactions are possible, such as a group of adenine bases binding to each other in a bulge, or the GNRA tetraloop that has a guanine–adenine base-pair[2].
An important structural feature of RNA that distinguishes it from DNA is the presence of a hydroxyl group at the 2' position of the ribose sugar. The presence of this functional group causes the helix to adopt the A-form geometry rather than the B-form most commonly observed in DNA. This results in a very deep and narrow major groove and a shallow and wide minor groove. A second consequence of the presence of the 2'-hydroxyl group is that in conformationally flexible regions of an RNA molecule (that is, not involved in formation of a double helix), it can chemically attack the adjacent phosphodiester bond to cleave the backbone.
Ribose is an aldopentose, that is a monosaccharide containing five carbon atoms that, in its open chain form, has an aldehyde functional group at one end. In the conventional numbering scheme for monosaccharides, the carbon atoms are numbered from C1' (in the aldehyde group) to C5'. The deoxyribose derivative, found in DNA, differs from ribose by having a hydrogen atom in place of the hydroxyl group in carbon C2'. Like many monosaccharides, ribose occurs in water as the linear form H-(C=O)-(CHOH)4-H and any of two ring forms: ribofuranose ("C3'-endo"), with a five-membered ring, and ribopyranose ("C2'-endo"), with a six-membered ring. The ribofuranose form is predominant in aqueous solution. The "D-" in the name D-ribose refers to the stereochemistry of the chiral carbon atom farthest away from the aldehyde group (C4'). In D-ribose, as in all D-sugars, this carbon atom has the same configuration as in D-glyceraldehyde. Ribose comprises the backbone of RNA, a biopolymer that is the basis of genetic transcription. It is related to deoxyribose, as found in DNA. Once phosphorylated, ribose can become a subunit of ATP, NADH, and several other compounds that are critical to metabolism[3][4].
RNA bases and its nucleotides
[edit | edit source]Adenine (A)
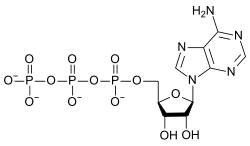
Adenine (A,) is a nucleobase (a purine derivative) with a variety of roles in biochemistry including cellular respiration, in the form of both the energy-rich adenosine triphosphate (ATP) and the cofactors nicotinamide adenine dinucleotide (NAD) and flavin adenine dinucleotide (FAD), and protein synthesis, as a chemical component of DNA and RNA. The shape of adenine is complementary to either thymine in DNA or uracil in RNA.
Cytosine (C)
Cytosine (C) is one of the four main bases found in DNA and RNA, along with adenine, guanine, and thymine (uracil in RNA). It is a pyrimidine derivative, with a heterocyclic aromatic ring and two substituents attached (an amine group at position 4 and a keto group at position 2). The nucleoside of cytosine is cytidine.
Guanine (G)
Guanine (G) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. With the formula C5H5N5O, guanine is a derivative of purine, consisting of a fused pyrimidine-imidazole ring system with conjugated double bonds.
Uracil (U)

Found in RNA, it base-pairs with adenine and replaces thymine during DNA transcription. Methylation of uracil produces thymine. It turns into thymine to protect the DNA and to improve the efficiency of DNA replication. Uracil can base-pair with any of the bases, depending on how the molecule arranges itself on the helix, but readily pairs with adenine because the methyl group is repelled into a fixed position. Uracil pairs with adenine through hydrogen bonding. Uracil is the hydrogen bond acceptor and can form two hydrogen bonds. Uracil can also bind with a ribose sugar to form the ribonucleoside uridine. When a phosphate attaches to uridine, uridine 5'-monophosphate is produced.[5]
Uracil undergoes amide-imidic acid tautomeric shifts because any nuclear instability the molecule may have from the lack of formal aromaticity is compensated by the cyclic-amidic stability. The amide tautomer is referred to as the lactam structure, while the imidic acid tautomer is referred to as the lactim structure. These tautomeric forms are predominant at pH 7. The lactam structure is the most common form of uracil.
Uracil also recycles itself to form nucleotides by undergoing a series of phosphoribosyltransferase reactions. Degradation of uracil produces the substrates aspartate, carbon dioxide, and ammonia.
- C4H4N2O2 → H3NCH2CH2COO- + NH4+ + CO2
Oxidative degradation of uracil produces urea and maleic acid in the presence of H2O2 and Fe2+ or in the presence of diatomic oxygen and Fe2+.
Uracil is a weak acid; the first site of ionization of uracil is not known.[6] The negative charge is placed on the oxygen anion and produces a pKa of less than or equal to 12. The basic pKa = -3.4, while the acidic pKa = 9.389. In the gas phase, uracil has 4 sites that are more acidic than water.[7]
Uracil is a common and naturally occurring pyrimidine derivative. Originally discovered in 1900, it was isolated by hydrolysis of yeast nuclein that was found in bovine thymus and spleen, herring sperm, and wheat germ. It is a planar, unsaturated compound that has the ability to absorb light.

Types of RNA
[edit | edit source]RNAs involved in protein synthesis
[edit | edit source]Messenger RNA
[edit | edit source]Messenger RNA (mRNA) is a molecule of RNA encoding a chemical "blueprint" for a protein product. mRNA is transcribed from a DNA template, and carries coding information to the sites of protein synthesis: the ribosomes. Here, the nucleic acid polymer is translated into a polymer of amino acids: a protein. In mRNA as in DNA, genetic information is encoded in the sequence of nucleotides arranged into codons consisting of three bases each. Each codon encodes for a specific amino acid, except the stop codons that terminate protein synthesis. This process requires two other types of RNA: transfer RNA (tRNA) mediates recognition of the codon and provides the corresponding amino acid, while ribosomal RNA (rRNA) is the central component of the ribosome's protein manufacturing machinery[8].

Ribosomal RNA
[edit | edit source]Ribosomal ribonucleic acid (rRNA) is the RNA component of the ribosome, the organelle that is the site of protein synthesis in all living cells. Ribosomal RNA provides a mechanism for decoding mRNA into amino acids and interacts with tRNAs during translation by providing peptidyl transferase activity. The tRNAs bring the necessary amino acids corresponding to the appropriate mRNA codon.
tRNA
[edit | edit source]Transfer RNA (tRNA) is a small RNA molecule (usually about 73-95 nucleotides) that transfers a specific active amino acid to a growing polypeptide chain at the ribosomal site of protein synthesis during translation. It has a 3' terminal site for amino acid attachment. This covalent linkage is catalyzed by an aminoacyl tRNA synthetase. It also contains a three base region called the anticodon that can base pair to the corresponding three base codon region on mRNA. Each type of tRNA molecule can be attached to only one type of amino acid, but because the genetic code contains multiple codons that specify the same amino acid, tRNA molecules bearing different anticodons may also carry the same amino acid[9][10].
RNAs involved in post-transcriptional
[edit | edit source]Small nuclear ribonucleic acid (snRNA)
[edit | edit source]Small nuclear ribonucleic acid (snRNA) is a class of small RNA molecules that are found within the nucleus of eukaryotic cells. They are transcribed by RNA polymerase II or RNA polymerase III and are involved in a variety of important processes such as RNA splicing (removal of introns from hnRNA), regulation of transcription factors (7SK RNA) or RNA polymerase II (B2 RNA), and maintaining the telomeres. They are always associated with specific proteins, and the complexes are referred to as small nuclear ribonucleoproteins (snRNP) or sometimes as snurps. These elements are rich in uridine content.
Small nucleolar RNAs (snoRNAs)
[edit | edit source]Small nucleolar RNAs (snoRNAs) are a class of small RNA molecules that primarily guide chemical modifications of other RNAs, mainly ribosomal RNAs, transfer RNAs and small nuclear RNAs. There are two main classes of snoRNA, the C/D box snoRNAs which are associated with methylation, and the H/ACA box snoRNAs which are associated with pseudouridylation. snoRNAs are commonly referred to as guide RNAs but should not be confused with the guide RNAs that direct RNA editing in trypanosomes.[11]
After transcription, nascent rRNA molecules (termed pre-rRNA) are required to undergo a series of processing steps in order to generate the mature rRNA molecule. Prior to cleavage by exo- and endonucleases the pre-rRNA undergoes a complex pattern of nucleoside modifications. These include methylations and pseudouridylations, guided by snoRNAs. Methylation is the attachment or substitution of a methyl group onto various substrates. The rRNA of humans contain approximately 115 methyl group modifications. The majority of these are 2'O-ribose-methylations ( where the methyl group is attached to the ribose group). Pseudouridylation is the conversion (isomerisation) of the nucleoside uridine to a different isomeric form pseudouridine(Ψ). Mature human rRNAs contain approximately 95 Ψ modifications. Each snoRNA molecule acts as a guide for only one (or two) individual modifications in a target RNA. In order to carry out modification, each snoRNA associates with at least four protein molecules in an RNA/protein complex referred to as a small nucleolar ribonucleoprotein (snoRNP). The proteins associated with each RNA depend on the type of snoRNA molecule (see snoRNA guide families below). The snoRNA molecule contains an antisense element (a stretch of 10-20 nucleotides) which are base complementary to the sequence surrounding the base (nucleotide) targeted for modification in the pre-RNA molecule. This enables the snoRNP to recognise and bind to the target RNA. Once the snoRNP has bound to the target site the associated proteins are in the correct physical location to catalyse the chemical modification of the target base[12].
Ribonuclease P (RNase P)
[edit | edit source]Ribonuclease P (RNase P) is a type of Ribonuclease which cleaves RNA. RNase P is unique from other RNases in that it is a ribozyme – a ribonucleic acid that acts as a catalyst in the same way that a protein based enzyme would. Its function is to cleave off an extra, or precursor, sequence of RNA on tRNA molecules
Telomerase RNA
[edit | edit source]Telomerase RNA component, also known as TERC, is an RNA gene found in eukaryotes, that is a component of telomerase used to extend telomeres. Telomerase RNAs differ greatly in sequence and structure between vertebrates, ciliates and yeasts, but they share a 5' pseudoknot structure close to the template sequence. The vertebrate telomerase RNAs have a 3' H/ACA snoRNA-like domain.
References
[edit | edit source]- ↑ http://en.wikipedia.org/w/index.php?title=RNA&oldid=424952312
- ↑ http://en.wikipedia.org/w/index.php?title=RNA&oldid=424952312
- ↑ http://en.wikipedia.org/w/index.php?title=RNA&oldid=424952312
- ↑ Higgs PG (2000). "RNA secondary structure: physical and computational aspects". Quarterly Reviews of Biophysics 33: 199–253.
- ↑ Horton, Robert H.; et al.Principles of Biochemistry. 3rd ed. Upper Saddle River, NJ: Prentice Hall, 2002.
- ↑ Zorbach, W.W. Synthetic Procedures in Nucleic Acid Chemistry: Physical and Physicochemical Aids in Determination of Structure. Vol 2. New York: Wiley-Interscience, 1973.
- ↑ Lee, J.K.; Kurinovich, Ma. J Am Soc Mass Spectrom.13(8), 2005, 985-95.
- ↑ http://en.wikipedia.org/w/index.php?title=Messenger_RNA&oldid=421869704
- ↑ http://en.wikipedia.org/w/index.php?title=Transfer_RNA&oldid=424698874
- ↑ Felsenfeld G, Cantoni G (1964). "Use of thermal denaturation studies to investigate the base sequence of yeast serine sRNA". Proc Natl Acad Sci USA 51 (5): 818–26.
- ↑ http://en.wikipedia.org/w/index.php?title=Small_nuclear_RNA&oldid=423606232
- ↑ http://en.wikipedia.org/w/index.php?title=Small_nuclear_RNA&oldid=423606232